PyHawkes implements a variety of Bayesian inference algorithms for discovering latent network structure given excitatory point process observations.
We provide a number of classes for building and fitting
mutually-excitatory point process (aka Hawkes process) models
with underlying network structure. Let's walk through a simple example
where we construct a discrete time model with three nodes, as in examples/discrete_demo.
The nodes are connected via an excitatory network such that a node's events
increase the likelihood of subsequent events on downstream nodes.
# Create a simple random network with K nodes a sparsity level of p
# Each event induces impulse responses of length dt_max on connected nodes
K = 3
p = 0.25
dt_max = 20
network = ErdosRenyiFixedSparsity(K, p)
true_model = DiscreteTimeNetworkHawkesModelSpikeAndSlab(K=K, dt_max=dt_max, network=network)
# Generate T time bins of events from the the model
# S is the TxK event count matrix, R is the TxK rate matrix
S,R = true_model.generate(T=1000)
true_model.plot()You should see something like this. Here, each event on node one adds
an impulse response on the rate of nodes two and three.
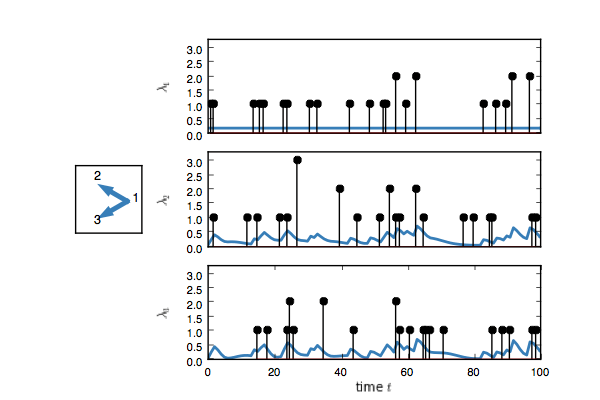
Now create a test model and try to infer the network given only the event counts.
# Create the test model, add the event count data, and plot
test_model = DiscreteTimeNetworkHawkesModelSpikeAndSlab(K=K, dt_max=dt_max, network=network)
test_model.add_data(S)
fig, handles = test_model.plot(color="#e41a1c")
# Run a Gibbs sampler
N_samples = 100
lps = []
for itr in xrange(N_samples):
test_model.resample_model()
lps.append(test_model.log_probability())
# Update plots
test_model.plot(handles=test_handles)If you enable interactive plotting, you should see something like this.
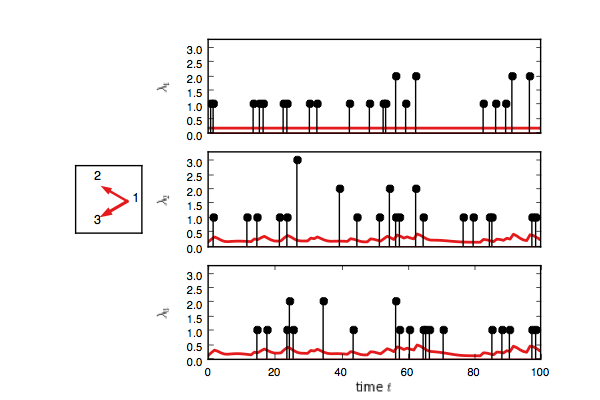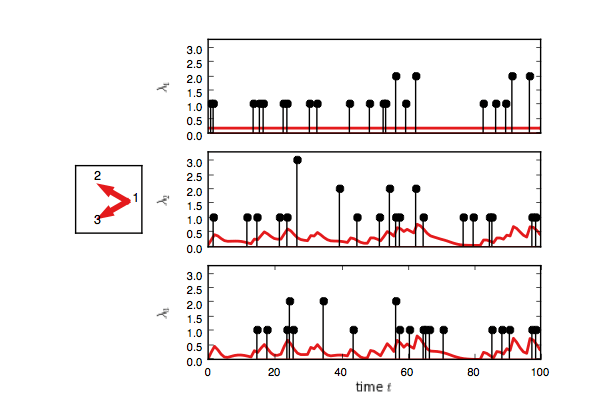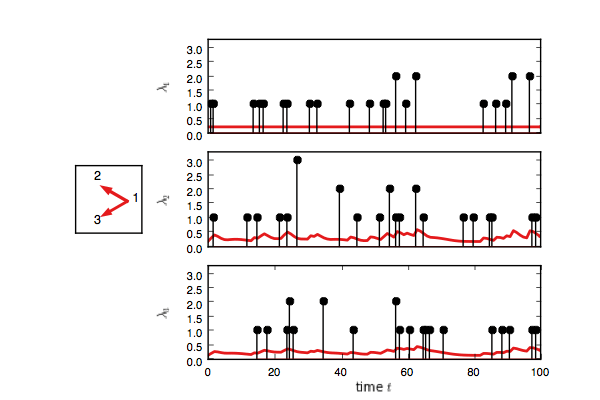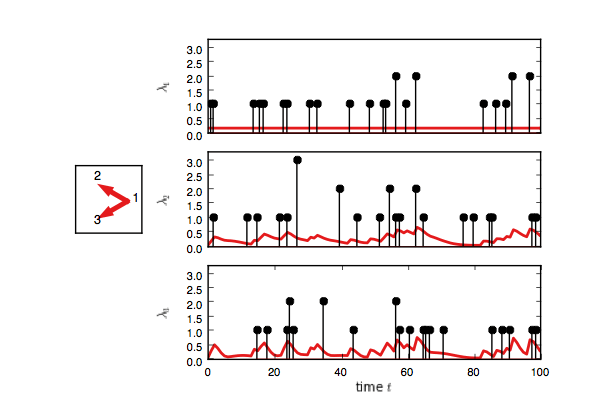
In addition to Gibbs sampling, we have implemented maximum a posteriori (MAP) estimation,
mean field variational Bayesian inference, and stochastic variational inference. To
see how those methods can be used, look in examples/inference.
To check out, run
git clone --recursive git@github.com:slinderman/pyhawkes.git
To compile the cython code, run
python setup.py build_ext --inplace
This codebase is considerably cleaner than the old CUDA version, and is still quite fast with the Cython+OMP extensions and joblib for parallel sampling of the adjacency matrix.
Complete details of this work can be found in:
Linderman, Scott W. and Adams, Ryan P. Discovering Latent Network Structure in Point Process Data. International Conference on Machine Learning (ICML), 2014.
and
Linderman, Scott W., and Adams, Ryan P. Scalable Bayesian Inference for Excitatory Point Process Networks. arXiv preprint arXiv:1507.03228, 2015.